In search of salamanders with environmental DNA
Biodiversity is in decline around the world; actions to conserve or recreate species-rich ecosystems are high on the global agenda. But how does one measure biodiversity? Researchers at IVL Swedish Environmental Research Institute are investigating this within a new research field: environmental DNA.
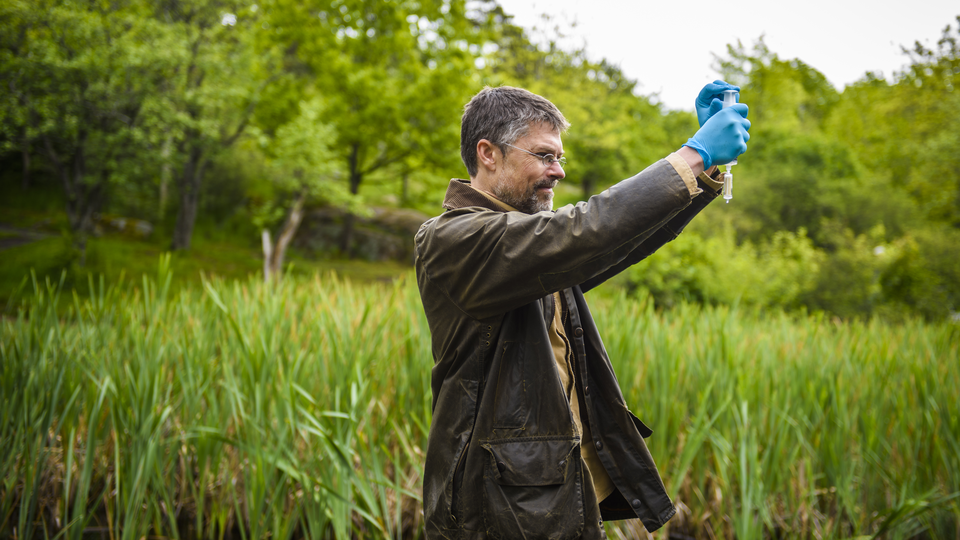
– OK Tage, time to contaminate the water!
Mats Töpel laughs, and searches his bag for a pair of blue plastic gloves, while Tage Vowles tries barehandedly to catch a small salamander in a frog pond at the top of Slottsskogen – a large park near central Gothenburg.
The situation very much resembles a crime scene – there are plastic gloves, sterile zip-lock bags, filters and various containers. And two meticulous experts, methodically carrying out their examination. But they are not looking for a perpetrator, but something very rare and difficult to catch. Both Swedish species of the water salamander are protected, and the larger water salamander is listed on the EU’s Habitats Directive, and has thus an even higher level of protection, e.g. that its breeding grounds are protected.
– Normally it is prohibited to disturb protected species, but the regulations on species protection have an exception whereby it is permitted to temporarily catch a small number for study purposes, which this certainly is, says Tage Vowles – who didn’t catch anything this time.
In any case, today’s sampling was less about the salamanders themselves. The focus was on testing the sampling equipment and recording an instruction film.
Environmental DNA – cheaper, faster, and increasingly relevant

– The technology behind environmental DNA is based on the technology that is used in the analysis of human DNA. All organisms leave traces of DNA. For instance, when a fish moves through the water, it constantly leaves behind traces of itself, such as dead skin cells, mucus or faeces. This organic material contains DNA molecules, and is called environmental DNA, explains Mats Töpel.
Biologist Mats Töpel began studying DNA molecules and sequences 15 years ago, and has seen how the technology has become cheaper, faster, and increasingly relevant in the environmental sector.
– Analysis of DNA, and the similar molecule RNA, in environmental sampling make it possible to conduct completely new types of studies. Examples include the analysis of Covid-19 in wastewater during the pandemic, or that you can look at genetic variation within species, which is difficult if not impossible using traditional methods. There is currently a huge amount of activity in this field.
Environmental DNA can even reveal what different organisms eat, or which plants a certain type of pollinator visits. Hidden biodiversity such as micro-organisms, unknown species or simply genetic variation within a species can be captured with environmental DNA.
A huge advantage compared to typical field inventories is that the actual sampling can be done by a layperson – with the right instructions – who can then submit the sample for analysis and interpretation.
Of particularly great interest to many operations today is the capability to see trends over time, for instance how species richness is affected by various measures. How much will biodiversity increase as a result of animals grazing on overgrown land? And how much will it decrease as a result of a certain type of development?

Does environmental DNA mean that field biology has lost relevance?
– No not at all, but it can be seen as a strong complement. For instance, the answers we get from a DNA test don’t say very much about how many individuals there are in a certain environment, or the age distribution in the population. For this reason, some studies are most easily done using traditional inventory methods. But if we are to tackle the environmental challenges we face, and manage the effects they have on biodiversity, we need access to inexpensive, new and reproducible methods, that include DNA sequencing of environmental samples.
Can we rely on the results of an environmental DNA analysis? Can’t they be misleading?
– It is very important to do the sampling in the right order, and with a relevant number of sampling points. Today we are taking four samples, which is plenty for being able to determine the presence of salamander. Because it’s such a small pond. Also, the concentration of DNA is greater in standing water than in moving water. And there is more DNA in the water in the summer, when most species breed or lay eggs.
And if you want an idea of the biodiversity in a section of forest?
– The larger the area, the more samples you need, but generally you get quite a good idea after the first sample. And there are different methods: you can search for a particular species, or you can sequence all the DNA in the sample by way of metagenomics.
While Mats and Tage filter their water samples, a moorhen floats past, through the reeds. A number of pigeons approach hopefully, over and over. And the occasional dog – and owner – pass by, curious about what is under way. People, animals and plants leave DNA traces – everywhere.
To minimize the risk of the molecules decomposing on the way to the lab, the samples are conserved in 99% alcohol, before being sent to IVL’s lab in Stockholm, where the DNA is sequenced. We already know that salamanders exist in the pond, but we will find traces of other rare species?